Projection
The swiftsimio.visualisation.projection sub-module provides an interface
to render SWIFT data projected to a grid. This takes your 3D data and projects
it down to 2D, such that if you request masses to be smoothed then these
functions return a surface density.
This effectively solves the equation:
\(\tilde{A}_i = \sum_j A_j W_{ij, 2D}\)
with \(\tilde{A}_i\) the smoothed quantity in pixel \(i\), and \(j\) all particles in the simulation, with \(W\) the 2D kernel. Here we use the Wendland-C2 kernel.
The primary function here is
swiftsimio.visualisation.projection.project_gas(), which allows you to
create a gas projection of any field. See the example below.
Example
from swiftsimio import load
from swiftsimio.visualisation.projection import project_gas
data = load("cosmo_volume_example.hdf5")
# This creates a grid that has units msun / Mpc^2, and can be transformed like
# any other cosmo_array
mass_map = project_gas(
data,
resolution=1024,
project="masses",
parallel=True,
periodic=True,
)
# Let's say we wish to save it as msun / kpc^2 (physical, not comoving),
from unyt import msun, kpc
from matplotlib.pyplot import imsave
from matplotlib.colors import LogNorm
# Normalize and save
imsave(
"gas_surface_dens_map.png",
LogNorm()(mass_map.to_physical_value(msun / kpc**2)),
cmap="viridis",
)
This basic demonstration creates a mass surface density map.
To create, for example, a projected temperature map, we need to remove the
surface density dependence (i.e. project_gas()
returns a surface temperature in units of K / kpc^2 and we just want K) by dividing out by
this:
from swiftsimio import load
from swiftsimio.visualisation.projection import project_gas
data = load("cosmo_volume_example.hdf5")
# First create a mass-weighted temperature dataset
data.gas.mass_weighted_temps = data.gas.masses * data.gas.temperatures
# Map in msun / mpc^2
mass_map = project_gas(
data,
resolution=1024,
project="masses",
parallel=True,
periodic=True,
)
# Map in msun * K / mpc^2
mass_weighted_temp_map = project_gas(
data,
resolution=1024,
project="mass_weighted_temps",
parallel=True,
periodic=True,
)
temp_map = mass_weighted_temp_map / mass_map
from unyt import K
from matplotlib.pyplot import imsave
from matplotlib.colors import LogNorm
# Normalize and save
imsave(
"temp_map.png",
LogNorm()(temp_map.to_physical_value(K)),
cmap="twilight",
)
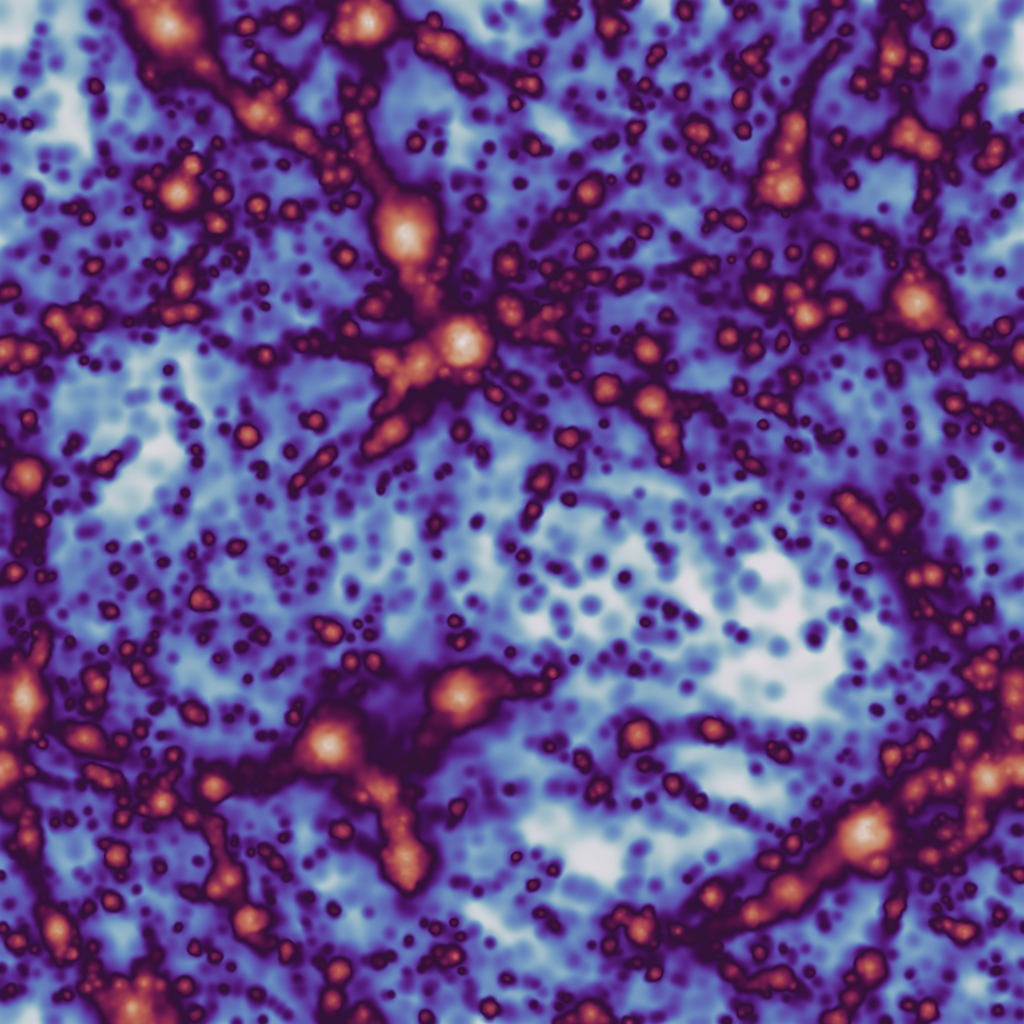
The output from this example, when used with the example data provided in the loading data section should look something like:
Backends
In certain cases, rather than just using this facility for visualisation, you
will wish that the values that are returned to be as well converged as
possible. For this, we provide several different backends. These are passed
as backend="str" to all of the projection visualisation functions, and
are available in the module
swiftsimio.visualisation.projection.projection_backends. The available
backends are as follows:
fast: The default backend - this is extremely fast, and provides very basic smoothing, with a return type of single precision floating point numbers.histogram: This backend provides zero smoothing, and acts in a similar way to thehistogram2d()function but with the same arguments asscatter.reference: The same backend asfastbut with two distinguishing features: all calculations are performed in double precision, and it will return early with a warning message if there are not enough pixels to fully resolve each kernel. Intended for developer usage, regular users should not use this mode.renormalised: The same asfast, but each kernel is evaluated twice and renormalised to ensure mass conservation within floating point precision. Returns single precision arrays.subsampled: This is the recommended mode for users who wish to have converged results even at low resolution. Each kernel is evaluated at least 32 times, with overlaps between pixels considered for every single particle. Returns in double precision.subsampled_extreme: The same assubsampled, but provides 64 kernel evaluations.gpu: The same asfastbut uses CUDA for faster computation on supported GPUs. The parallel implementation is the same function as the non-parallel.
Example:
from swiftsimio import load
from swiftsimio.visualisation.projection import project_gas
data = load("cosmo_volume_example.hdf5")
subsampled_array = project_gas(
data,
resolution=1024,
project="entropies",
parallel=True,
backend="subsampled",
periodic=True,
)
This will likely look very similar to the image that you make with the default
backend="fast", but will have a well-converged distribution at any resolution
level.
Periodic boundaries
Cosmological simulations and many other simulations use periodic boundary conditions. This has implications for the particles at the edge of the simulation box: they can contribute to pixels on multiple sides of the image. If this effect is not taken into account, then the pixels close to the edge will have values that are too low because of missing contributions.
All visualisation functions by default assume a periodic box. Rather than
simply projecting each individual particle once, four additional periodic copies
of each particle are also projected. Most copies will project outside the valid
pixel range, but the copies that do not ensure that pixels close to the edge
receive all necessary contributions. Thanks to numba optimisations, the overhead
of these additional copies is relatively small.
There are some caveats with this approach. If you try to visualise a subset of the particles in the box (e.g. using a mask), then only periodic copies of particles in this subset will be used. If the subset does not include particles on the other side of the periodic boundary, then these will still be missing from the projection. The same is true if you visualise a region of the box. The periodic boundary wrapping is also not compatible with rotations (see below) and should therefore not be used together with a rotation.
Rotations
Sometimes you will need to visualise a galaxy from a different perspective.
The swiftsimio.visualisation.rotation sub-module provides routines to
generate rotation matrices corresponding to vectors, which can then be
provided to the rotation_matrix argument of
project_gas() (and
project_gas_pixel_grid()). You will also need
to supply the rotation_center argument, as the rotation takes place around this given
point. The example code below loads a snapshot, and a halo catalogue, and
creates an edge-on and face-on projection using the integration in
velociraptor. More information on possible integrations with this library
is shown in the velociraptor section.
from swiftsimio import load, mask, cosmo_array
from velociraptor import load as load_catalogue
from swiftsimio.visualisation.rotation import rotation_matrix_from_vector
from swiftsimio.visualisation.projection import project_gas
import unyt
import numpy as np
# Radius around which to load data, we will visualise half of this
size = 1000 * unyt.kpc
snapshot_filename = "cosmo_volume_example.hdf5"
catalogue_filename = "cosmo_volume_example.properties"
catalogue = load_catalogue(catalogue_filename)
# Which halo should we visualise?
halo = 0
x = catalogue.positions.xcmbp[halo]
y = catalogue.positions.ycmbp[halo]
z = catalogue.positions.zcmbp[halo]
lx = catalogue.angular_momentum.lx[halo]
ly = catalogue.angular_momentum.ly[halo]
lz = catalogue.angular_momentum.lz[halo]
# The angular momentum vector will point perpendicular to the galaxy disk.
# If your simulation contains stars, use lx_star
angular_momentum_vector = cosmo_array([lx, ly, lz])
angular_momentum_vector /= np.linalg.norm(angular_momentum_vector)
face_on_rotation_matrix = rotation_matrix_from_vector(angular_momentum_vector)
edge_on_rotation_matrix = rotation_matrix_from_vector(
angular_momentum_vector, axis="y"
)
data_mask = mask(snapshot_filename)
region = cosmo_array(
[
[x - size, x + size],
[y - size, y + size],
[z - size, z + size],
],
x.units,
comoving=True,
scale_factor=data_mask.metadata.a,
scale_exponent=1,
)
visualise_region = cosmo_array(
[
x - 0.5 * size,
x + 0.5 * size,
y - 0.5 * size,
y + 0.5 * size,
],
comoving=True,
scale_factor=data_mask.metadata.a,
scale_exponent=1,
)
data_mask.constrain_spatial(region)
data = load(snapshot_filename, mask=data_mask)
# Use project_gas_pixel_grid to generate projected images
common_arguments = dict(
data=data,
resolution=512,
parallel=True,
region=visualise_region,
periodic=False, # disable periodic boundaries when using rotations
)
un_rotated = project_gas(**common_arguments)
rotation_center = cosmo_array(
[x, y, z], comoving=True, scale_factor=data_mask.metadata.a, scale_exponent=1
)
face_on = project_gas(
**common_arguments,
rotation_center=rotation_center,
rotation_matrix=face_on_rotation_matrix,
)
edge_on = project_gas(
**common_arguments,
rotation_center=rotation_center,
rotation_matrix=edge_on_rotation_matrix,
)
Using this with the provided example data will just show blobs due to its low resolution
nature. Using one of the EAGLE volumes (examples/EAGLE_ICs) will produce much nicer
galaxies, but that data is too large to provide as an example in this tutorial.
You can also provide an extra two values, the z min and max, as part of the
region parameter. This may have some slight performance impact, so it is
generally advised that you do this on sub-loaded volumes only.
Other particle types
Other particle types are able to be visualised through the use of the
swiftsimio.visualisation.projection.project_pixel_grid() function.
To use this feature for particle types that do not have smoothing lengths, you
will need to generate them, as in the example below where we create a
mass density map for dark matter. We provide a utility to do this through
generate_smoothing_lengths().
from swiftsimio import load
from swiftsimio.visualisation.projection import project_pixel_grid
from swiftsimio.visualisation.smoothing_length import generate_smoothing_lengths
data = load("cosmo_volume_example.hdf5")
# Generate smoothing lengths for the dark matter
data.dark_matter.smoothing_length = generate_smoothing_lengths(
data.dark_matter.coordinates,
data.metadata.boxsize,
kernel_gamma=1.8,
neighbours=57,
speedup_fac=2,
dimension=3,
)
# Project the dark matter mass
dm_mass = project_pixel_grid(
# Note here that we pass in the dark matter dataset not the whole
# data object, to specify what particle type we wish to visualise
data=data.dark_matter,
resolution=1024,
project="masses",
parallel=True,
region=None,
periodic=True,
)
from matplotlib.pyplot import imsave
from matplotlib.colors import LogNorm
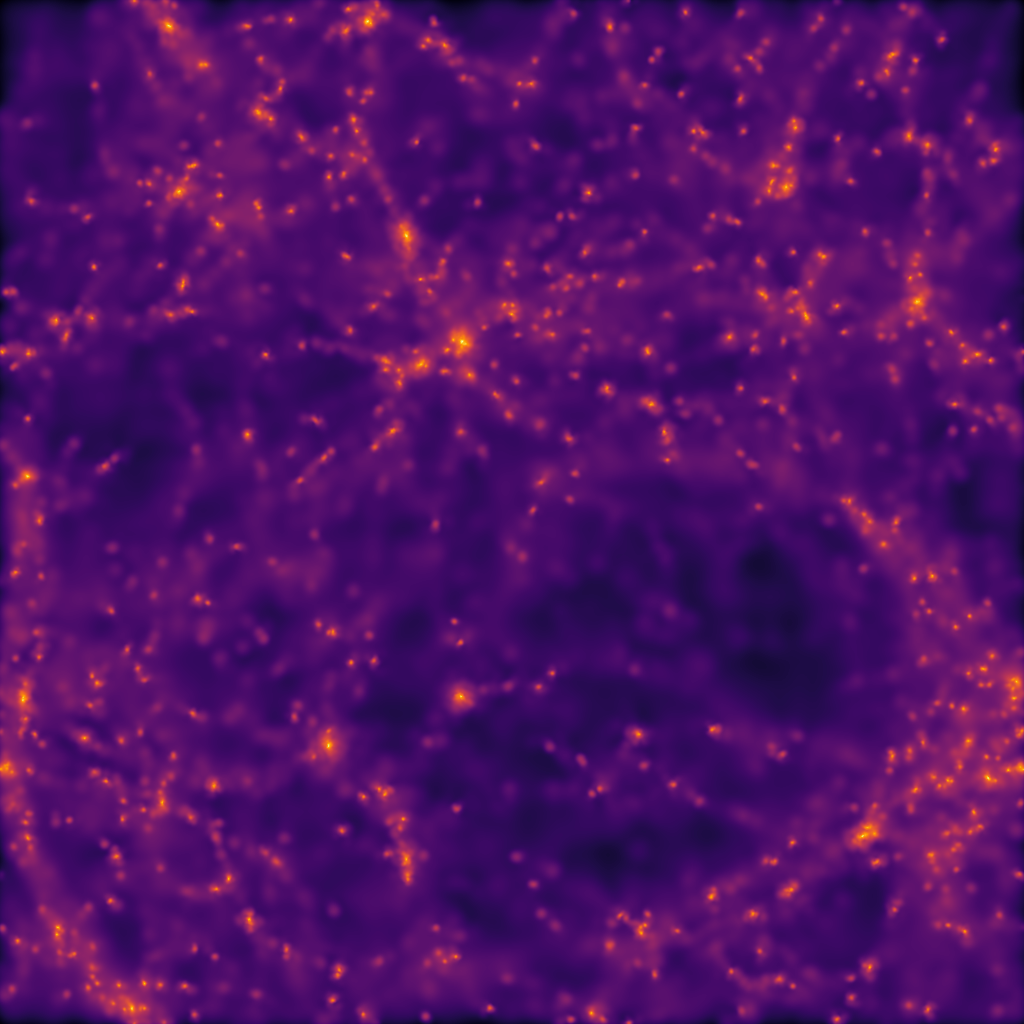
# Everyone knows that dark matter is purple
imsave("dm_mass_map.png", LogNorm()(dm_mass), cmap="inferno")
The output from this example, when used with the example data provided in the loading data section should look something like:

Lower-level API
The lower-level API for projections allows for any general positions, smoothing lengths, and smoothed quantities, to generate a pixel grid that represents the smoothed version of the data.
This API is available through
swiftsimio.visualisation.projection_backends.backends and
swiftsimio.visualisation.projection_backends.backends_parallel for parallel
implementations. The parallel versions use significantly more memory as they allocate
a thread-local image array for each thread, summing them in the end. Here we
will only describe the fast variant, but they behave in the exact same way.
The default backend used by swiftsimio is fast. To use the others (e.g.
histogram, subsampled, gpu, etc.), you can select them
manually from the module, or by using the backends and backends_parallel
dictionaries in swiftsimio.visualisation.projection_backends.
To use these functions, you will need:
x-positions of all of your particles,
x.y-positions of all of your particles,
y.A quantity which you wish to smooth for all particles, such as their mass,
m.Smoothing lengths for all particles,
h.The resolution you wish to make your square image at,
res.
Optionally, you will also need:
+ the size of the simulation box in x and y, box_x and box_y.
The key here is that only particles in the domain [0, 1] in x, and [0, 1] in y
will be visible in the image. You may have particles outside of this range;
they will not crash the code, and may even contribute to the image if their
smoothing lengths overlap with [0, 1]. You will need to re-scale your data
such that it lives within this range. You should also pass raw numpy arrays (not
cosmo_array or unyt_array, the
inputs are dimensionless). Then you may use the functions as follows:
from swiftsimio.visualisation.projection_backends import backends
scatter = backends["fast"]
# Using the variable names from above
out = scatter(x=x, y=y, h=h, m=m, res=res)
out will be a 2D ndarray grid of shape [res, res]. You will need
to re-scale this back to your original dimensions to get it in the correct units,
and do not forget that it now represents the smoothed quantity per surface area.
If the optional arguments box_x and box_y are provided, they should
contain the simulation box size in the same re-scaled coordinates as x and
y. The projection backend will then correctly apply periodic boundary
wrapping. If box_x and box_y are not provided or set to 0, no
periodic boundaries are applied.